EpiDirect® MLH1 Methylation qPCR Assay, CE IVD, IVDR
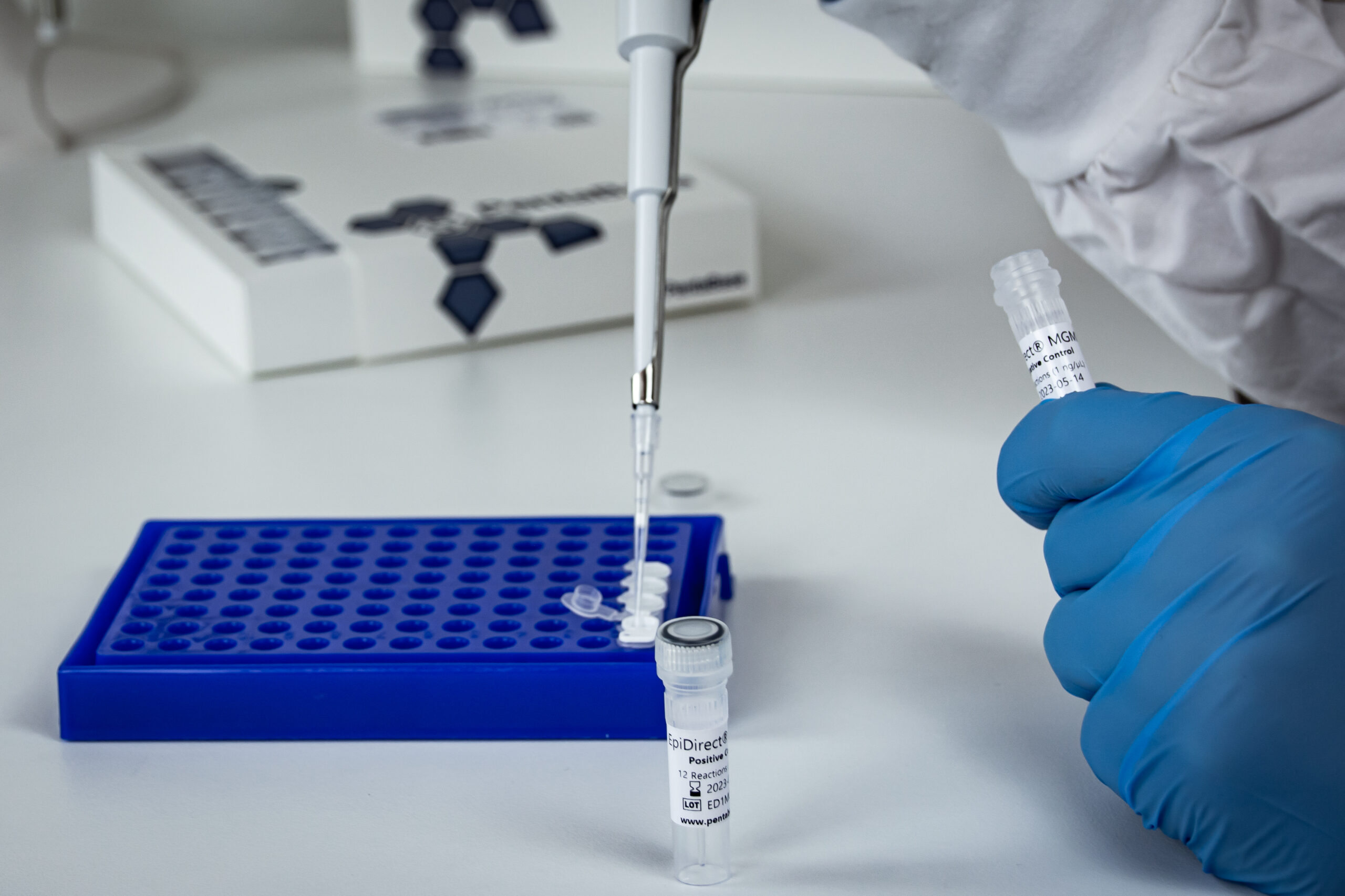
The EpiDirect® MLH1 Methylation qPCR Assay represents a new and innovative approach allowing for direct quantification of MLH1 promoter methylation by qPCR without prior DNA conversion.
The EpiDirect® technology is based on our unique INA® chemistry and includes an EpiPrimer™ oligonucleotide that specifically binds and primes amplification of methylated target DNA.
The assay is able to quantify MLH1 promoter methylation using standard real-time PCR instruments* in less than two hours with as little as 0.5 ng of unconverted input DNA.
- Direct DNA methylation analysis by qPCR without DNA pretreatment
- Standard qPCR instrument procedure
- No need for prior conversion of DNA
- Results in less than 2 hours
- Minimal hands-on time
- 0.5-50 ng input DNA
- CE IVD
- IVDR certified^
*Validated on CFX96/CFX Opus 96 (Bio-Rad), QuantStudio™5 (ABI) and BaseTyper™ 48.4 (PentaBase) Real-Time PCR Systems
^Conformity assessment by Notified Body Number 2797
Single Registration Number: DK-MF-000023157
Product Details
Reference Number:
8401
Product Name:
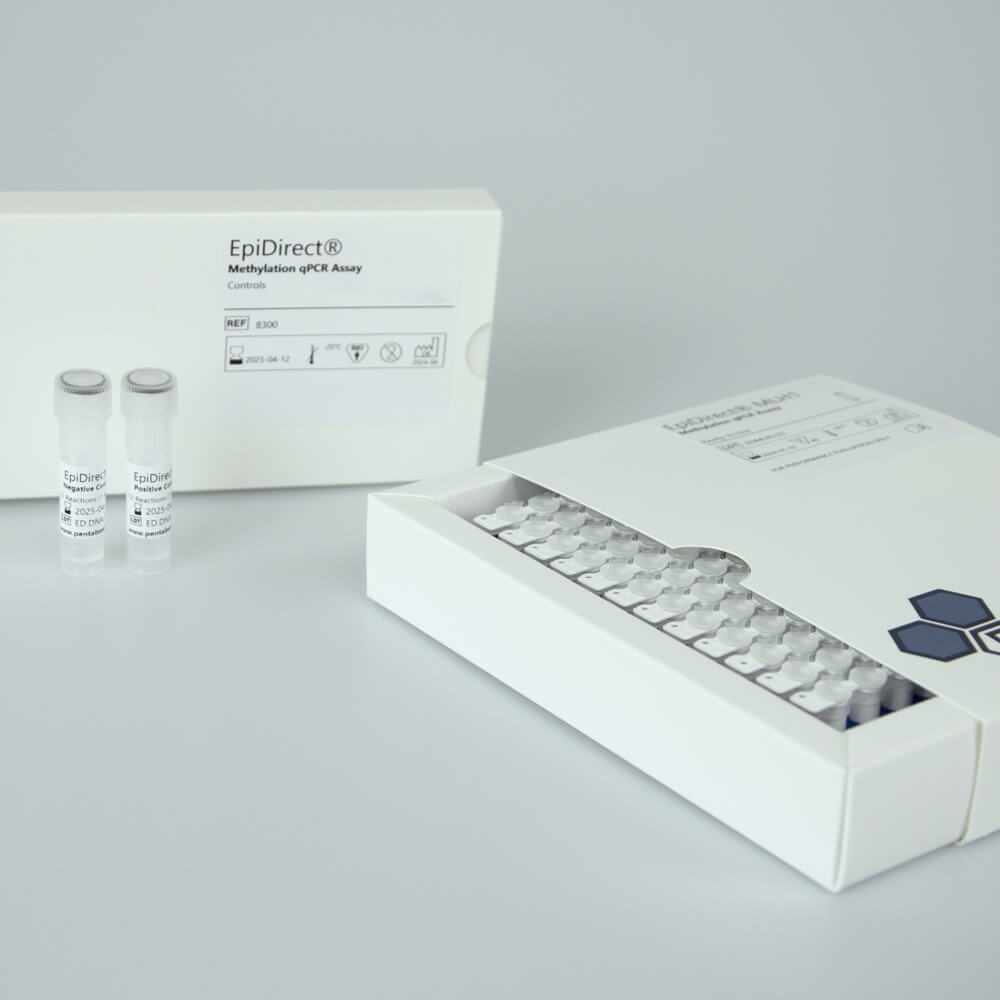
EpiDirect® MLH1 Methylation qPCR Assay
Product Description:
Includes PCR master mix and oligonucleotide mix for quantification of MLH1 promoter methylation, Positive Control, Negative Control and Standard Curve Controls
Format:
Ready-to-Use
Reactions:
48
Intended Use
EpiDirect® MLH1 Methylation qPCR Assay is a real-time polymerase chain reaction (PCR) assay intended to detect and quantify the methylation status of the human mismatch repair gene MutL Homolog1 (MLH1) promoter to aid in the diagnosis of Lynch Syndrome. The assay can be used qualitatively by applying the included positive control or quantitatively by applying the included standard curve. The assay is intended for use with purified human genomic DNA extracted from formalin-fixed paraffin-embedded tumor tissue from patients with colorectal or endometrial carcinoma. Without further pre-treatment, the samples can be analysed directly by semi-automatic real-time PCR systems
The assay is intended for use by qualified laboratory personnel specifically instructed and trained in the techniques of real-time PCR, as well as proficient in handling biological samples
Product Specifications
Procedure:
Real-time PCR-based
Validated Real-Time PCR instruments:
BaseTyper™ 48.4Quiet HRM (PentaBase Ref. No. 754), CFX96 and CFX Opus 96 (Bio-Rad) and QuantStudio™ 5 Real-Time PCR Systems
PentaBase technologies:
EpiPrimer™, HydrolEasy® Probe, SuPrimer™
Genomic target:
CpG sites 1-4 in MLH1 promoter Region C (-248 to -178)
Sample types:
Formalin-fixed paraffin-embedded (FFPE) or fresh-frozen solid colorectal and endometrial cancer biopsies
Sample tumor cell percentage:
At least 20%
Sample DNA extraction methods:
Relevant commercially avaliable solid biospy/FFPE DNA extraction kits
Sample input:
0.1-10 ng/µL of purified DNA directly from the sample that has not been otherwise chemically manipulated
PCR run time:
Less than 2 hours
Instructions For Use
For the latest version of the Instructions For Use (IFU) – Click here –
Publications
A qPCR technology for direct quantification of methylation in untreated DNA
Bendixen et al. – Nature Communications – 2023
Related files
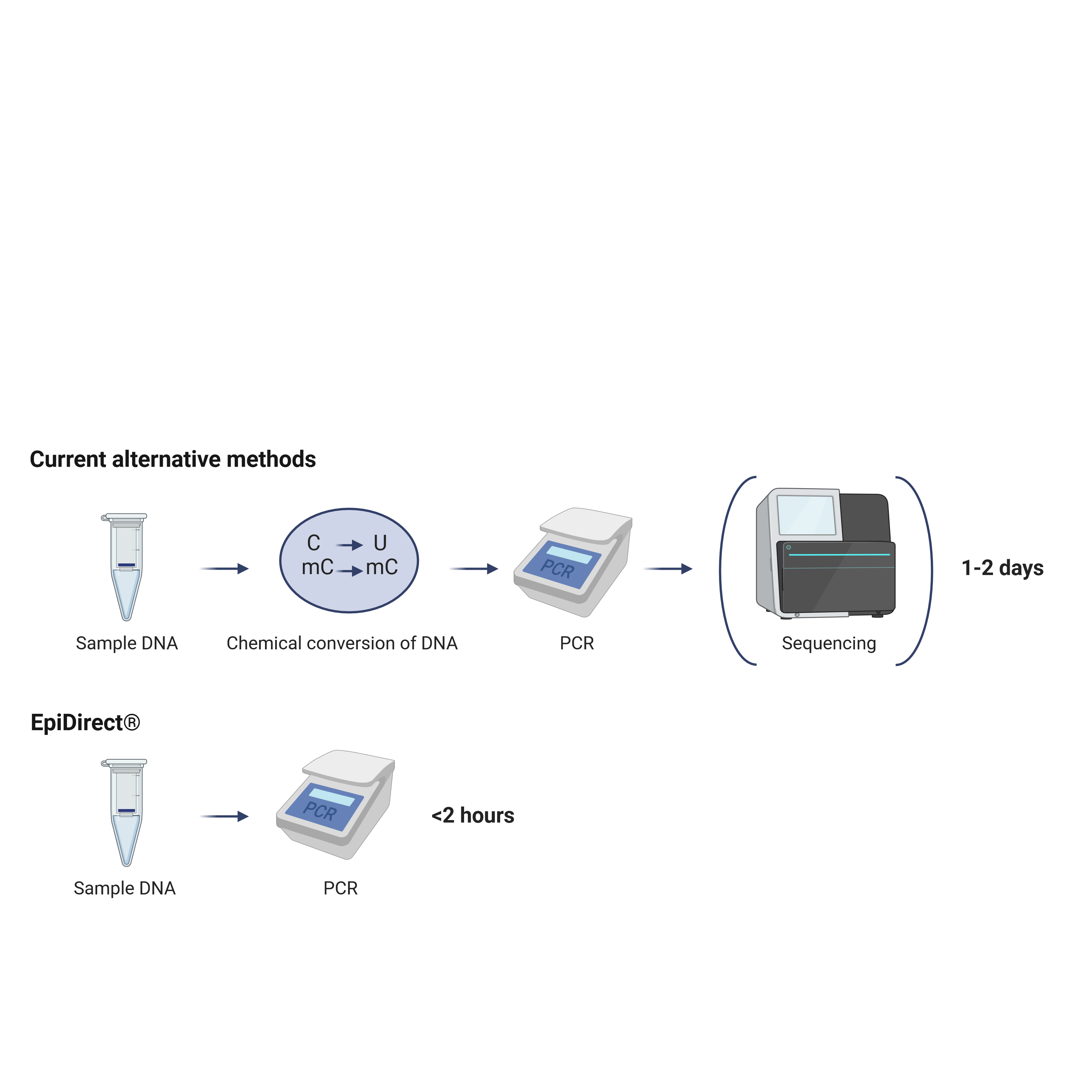
A Revolutionary Approach to DNA Methylation Testing
Current methods for evaluating DNA methylation are relying on sodium bisulfite or other chemical or enzymatical conversion of cytosines. However, variability in conversion rates as well as DNA degradation resulting from chemical treatment may lead to false or in-conclusive results. Especially bisulfite treatments have been shown to lead to a biased degradation of unmethylated DNA and consequently an overestimation of the methylation frequency.
In contrast, the EpiDirect® technology allows for direct quantification of DNA methylation by PCR without any prior chemical or enzymatic treatment. This revolutionizing technology not only eliminates artefacts related to incomplete conversion of cytosines and DNA degradation but also vastly reduces hands-on-time and time to result.
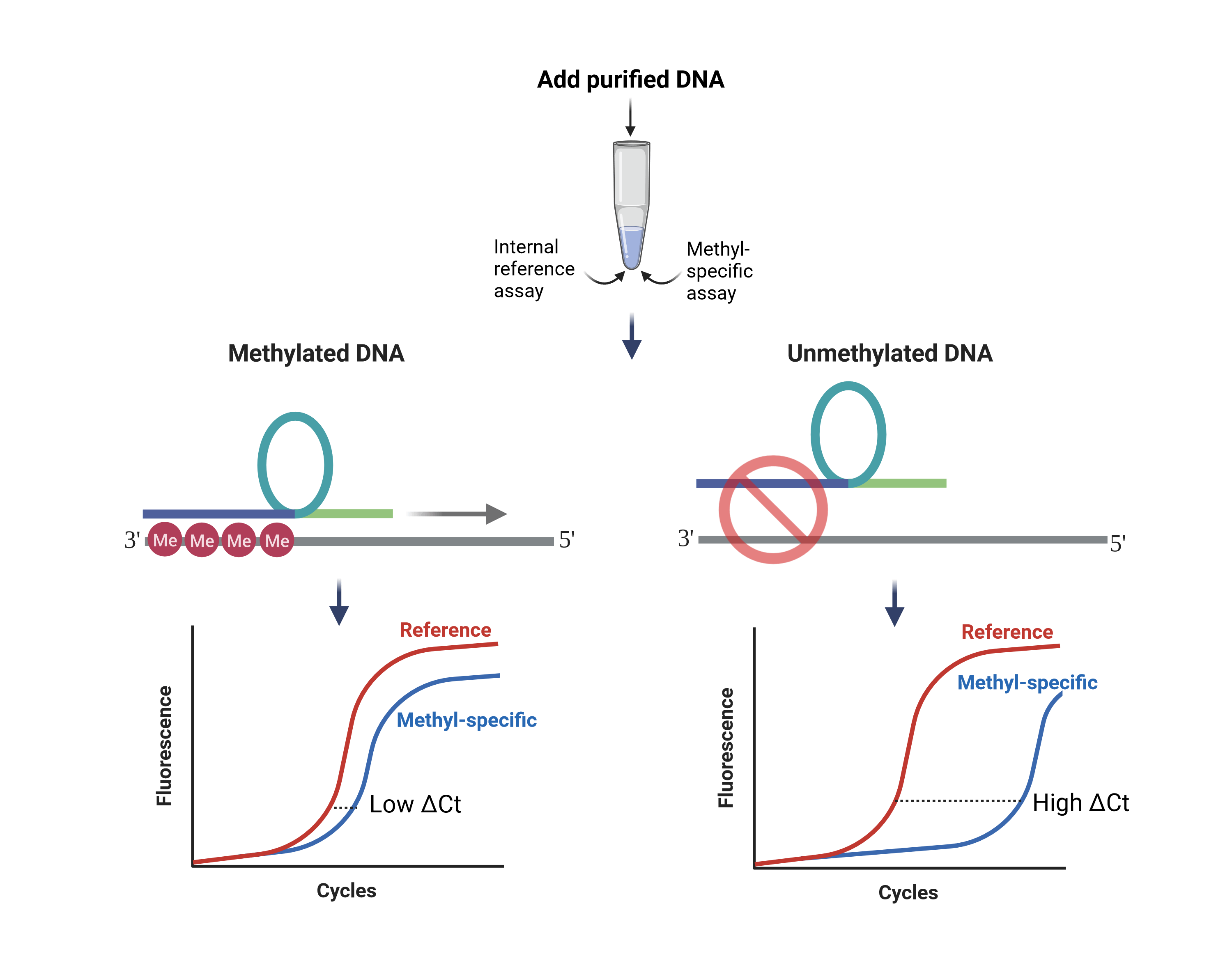
The EpiDirect® Technology
The EpiDirect® technology is based on a unique DNA analogue chemistry known as intercalating pseudo-nucleotide (IPN). IPN-containing oligonucleotides, also known as intercalating nucleic acids (INAs®), show highly improved target affinity and specificity compared to standard chemistry oligonucleotides.
The EpiDirect® technology utilizes a type of INA® known as an EpiPrimer™ that selectively binds to and primes amplification of methylated DNA. The EpiPrimer™ consists of 3 parts: a) an IPN-containing anchor sequence that binds to the methylated target region, b) a starter sequence that primes DNA replication, and c) a loop sequence that is part of the subsequent methyl-unspecific priming. Thus, the anchor and starter sequence prime the initial replication of the methylated DNA while the loop and starter sequence combined work as a primer in the remaining PCR cycles.